We then pipe the output to bcftools, which does our SNP calling based on those likelihoods. Samtools calculates the genotype likelihoods. Variant Calling using Samtools (Mpileup + bcftools) ¶
#WHAT IS BAM FILE FORMAT SOFTWARE#
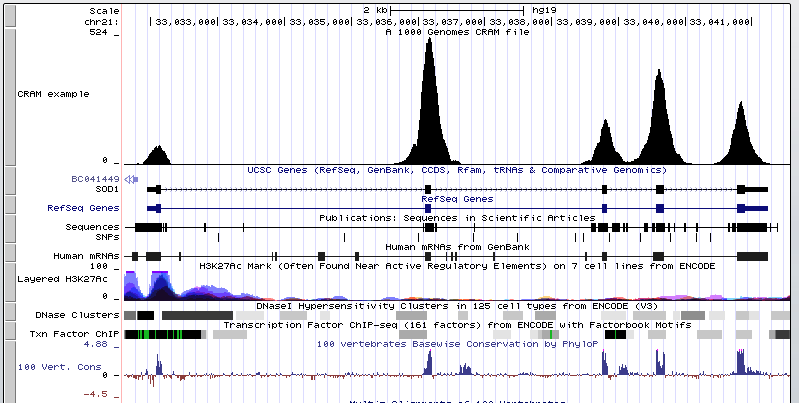
There a number of software available for variant calling some of which are as follows: What should be noted about variants is that they are rare events and homozygous variants are even rarer than heterozygous events. Variant calling will look at how many bases out of the total number of bases is different to the reference at any position. The data is converted into positional information of the reference with the read counts that have piled up under each position.

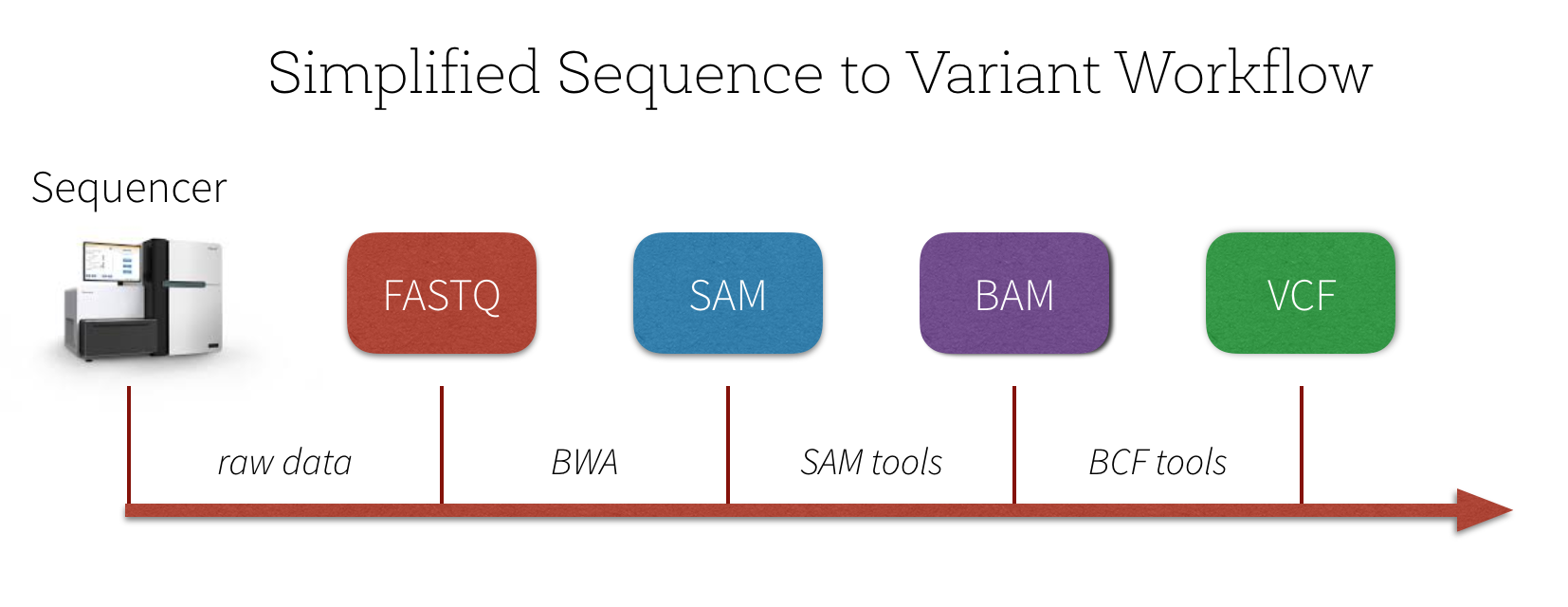
The number of reads that stack up on each other is called read coverage. Sample DNA -> Sequencing -> Read alignment -> BAM file of aligned reads against reference genome -> Genotyper -> Variant list The result of variant calling is a list of probable variants. The remaining question is, given that our variant calling process calls a variant, does that mean that there is truly a variant in this locus and also given that the variant caller doesn’t detect a variant in a position does that mean there is no variant in that position. The question one would be asking is what possible genotypes would be possible for a sample. Variant callers estimate the probability of a particular genotype given the observed data. In haploid organisms variant calling and genotyping are equivalent whereas the same rule does not apply to other organisms. Variant calling is concerned with whether there is evidence of variant in a particular locus whereas genotyping talks about what the sets of alleles in that locus are and their frequencies. Large Chromosome rearrangements-Structural variations (SV).Small insertions and deletions (INDELs).What sort of variation could we find in the DNA sequencing?
In Craig Venter’s genome 4.1 million DNA variants were reported. The aim of variation detection is to detect how many bases out of the total are different to a reference genome. Introduction to Variant detection ¶ Background ¶Ī variant is something that is different from a standard or type. Molecular Dynamics - Building input files, visualising the trajectory Molecular Dynamics - Introduction to cluster computing Identifying proteins from mass spectrometry data RNAseq differential expression tool comparision (Galaxy) Introduction to Metabarcoding using Qiime2 Hybrid genome assembly - Nanopore and Illumina Introduction to de novo genome assembly for Illumina readsĭe novo assembly of Illumina reads using Velvet (Galaxy)ĭe novo assembly of Illumina reads using Spades (Galaxy)
Introduction to de novo assembly with Velvet Variant Calling using GATK-Unified GenotyperĮvaluation of detected variants using Variant Eval Variant Calling using Samtools (Mpileup + bcftools) Common Workflow Language for Bioinformatics
